
If generated and provided by the ABIF or SCF file, then it will show the primary calls. What is shown will depend on how they were generated. Primary, Secondary or both basecalls can be shown if they are contained in the sangerseq object provided. It was acquired by MacVector, Inc on 1 January 2007. This function outputs a chromatogram to the current device or to a PDF file (filename is not NULL). Oxford Molecular was merged into Accelrys in 2001. It was acquired by Kodak, and subsequently Oxford Molecular in 1996. MacVector was originally developed by IBI in 1994. MacVector has a contig assembly plugin called Assembler that uses phred, phrap, Bowtie, SPAdes, Velvet and cross match.Īs of version 13.0.1 MacVector uses Sparkle for updating between releases. MacVector scans nucleic acid sequences for restriction enzymes and displays the cut sites in the Map view automatically.
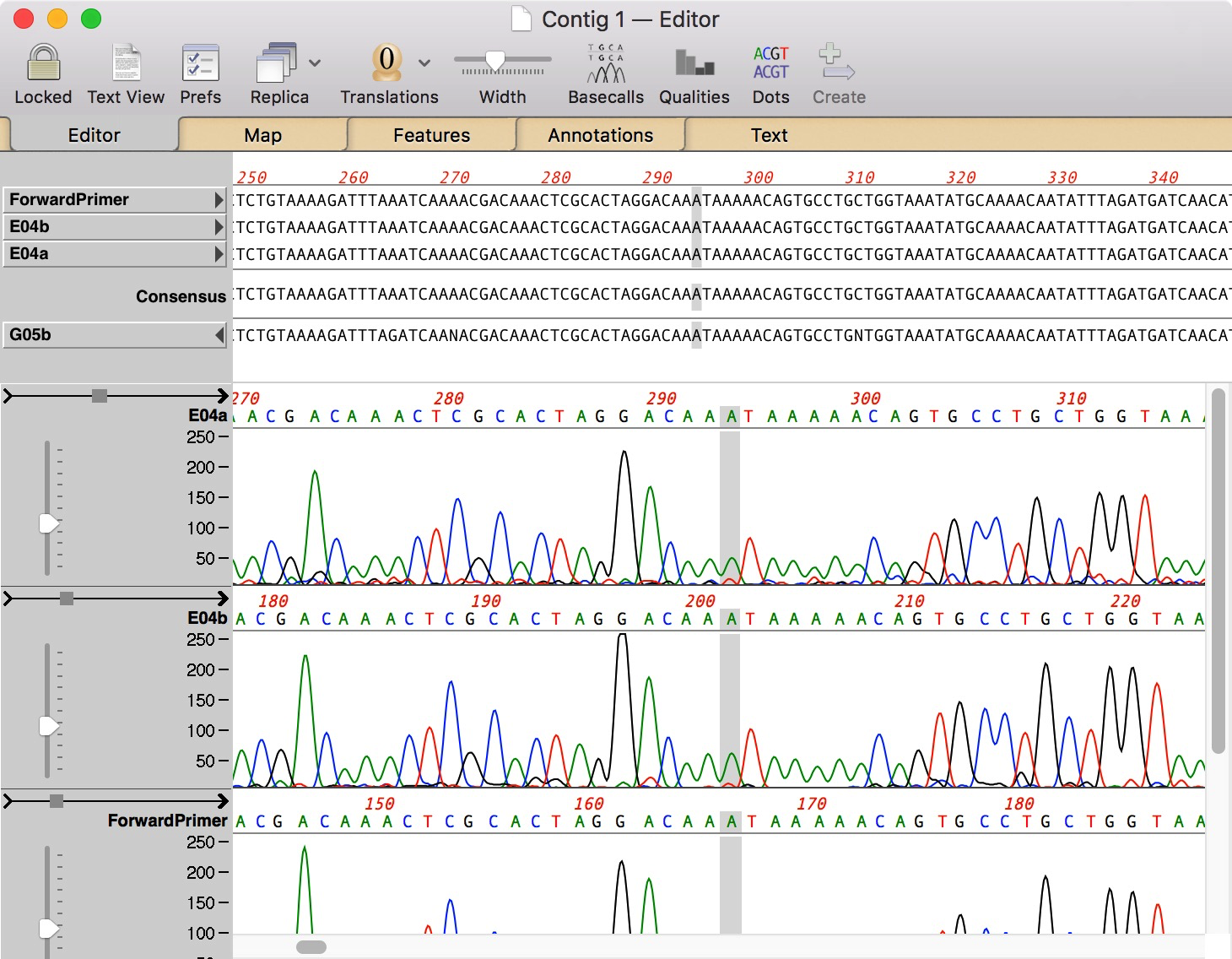

You will have to do it manually in DNA Sequence Assembler. If the QV data is missing the program cannot automatically cut the ends. The program uses the quality value (QV) data inside the chromatogram to automatically detect low quality regions to cut.and the cut-offs for acceptable scores were 0.5 and 0.2, respectively. Basically in this case the program will only remove the low quality ends and save the file back as SCF. DNA sequence analyses were performed using the MacVector 9 DNA sequence analysis. Don't worry if your chromatograms are already in SCF format.The output will the written in the same folder.'Convert all' converts all chromatograms in that folder. The 'Convert' button converts only the selected chromatogram.In the 'Convert panel' check the 'Remove low quality ends' and convert all your chromatograms to SCF.In 'Low quality end trimming' panel choose how much you want to cut from your chromatograms.Navigate to the folder where your ABI/SCF chromatograms are located.You can automatically trim low quality ends to all chromatograms in a folder. How to automatically trim low quality ends in all chromatograms DNA Chromatogram Explorer-Automatically trim low quality ends in chromatograms
